To test how intricate of a shape the wavelet transform of the brain-skull interface could be, I first started by tracing the edge using MultiTracer. Through that program I was able to export by traced points unto a UCF file so that I could work with the points directly. The following shows the brain slice I used with red outlining where I traced, followed by a plot of just the points I traced in red.


Once the tracing had been found separated the x and y points in the data to be modeled separately and also resampled them so that the total number would be a power of two to help fit in with the level framework in multiresolution analysis. Thus I had a total of 2^13 points. At first I tried to use the Cubic B-spline wavelets that I had tried before to model the ellipses but ran into a few problems. Once the function or in this case the data to be modeled became very intricate and complex the linear system I needed to solve became more troublesome to work with and the pseudo inverse I was using would not suffice to solve the system. To remedy this I tried to make the number of wavelets I had match the number of samples I was using to discretize the wavelets in hopes to make a square matrix that would easily be invertible. But the algorithm that I was using was using huge matrix operations and was thus taking too long to get to levels that would allow for the amount of detail that I needed.
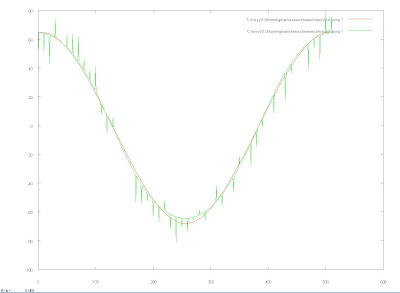
I decided to try using the Daubechies Wavelets since they seemed to be good for encoding polynomials. There also were algorithms created that took advantage of the fast wavelet transform that would enable me to avoid inverting the matrix and instead just calculate the coefficients at a high level and then filter them to lower levels, which proved to be much more efficient. The following graphs demonstrate my results. The first one is the tracing in green modeled by wavelets in red using level j=5 with 2^6 coefficients.

These next images represent the data in green modeled by the wavelets in red at levels j=8 and j=9 with 2^9 and 2^10 coefficients respectively. As the level increases the level of detail that the wavelet transform is able to capture also increases.


The following image represents level j=9 where I used only half the wavelets and the results show that the wavelet model of the data shown in green only covers half of the data. This demonstrated the ability to zero in on certain locations using the wavelets.


 Then I ran the optimizer using the optimal parameters that I found from the tracing and the following movie shows the results. The movie illustrates how the fit on the right side of the brain is closer then with just the ellipse but on the left the coefficients seem to be standing still, which may be due to some bugs in the optimizer that I have to iron out.
Then I ran the optimizer using the optimal parameters that I found from the tracing and the following movie shows the results. The movie illustrates how the fit on the right side of the brain is closer then with just the ellipse but on the left the coefficients seem to be standing still, which may be due to some bugs in the optimizer that I have to iron out.